Genetic Evidence for HERV-K102 Deletion From the Orthologous Position on Chromosome 1q22 in the Extinct Hominins: Neanderthals and Denisovans
Help from a newly minted Nobel Laureate.
Dr. Svante Paabo who just won the Nobel Prize in Medicine in 2022, allowed me the opportunity to scan his extinct hominin databases a number of years ago for HERV-K102 status at the orthologous 1q22 position. The hypothesis was that Homo sapiens might have had a survival advantage during the encounters with Neanderthal or Denisovans due to preferential retention of HERV-K102, a very powerful INNATE virus anti-virus mechanism to protect humans against emerging viruses, epidemics and pandemics.
The inquiry did not disappoint as discussed below.
Phylogenetic Evidence Supporting HERV-K102 in Defending Against RNA Viruses
Accumulating phylogenetic evidence is consistent with a potential role of HERV-K HML-2 in limiting invasion by orthoretroviruses (Magiorkinis et al., 2015).
Ancestral HML-2 elements emerged about 10.3 million years ago (mya) (Subramanian et al., 2011). There has been a striking decline of insertions of endogenous retroviruses (ERVs) in the last 10 million years (My) in the genomes of all sequenced hominids (great apes and gibbons), but not in old world monkeys (baboons and macaques), particularly regarding HERV-H (Magiorkinis et al., 2015). HERV-H makes up 88% of all the ERV integrations into the human genome in the last 30 My and became extinct over the past 10 My. HERV-H is a gammaretrovirus, which integrated around 45 to 60 mya and has about 962 copies in the human genome (Chuong et al., 2016). HERV-K, with 10 groups in the clade, only one of which is HML-2, on the other hand, entered the genome of ancestral catarrhines about 32 to 44 mya, after the split from New World monkeys and before the split of hominids from the Old World monkeys (Kim & Han, 2015). The sister lineages of HERV-K in most other catarrhines appear to have become extinct.
Most remarkably, the HERV-K HML-2 group in humans is the only HERV-K that has continued to replicate since the origin of the catarrhines (Magiorkinis et al., 2015). HERV-K102 is a member of the bioactive HERV-K HML-2 group and appears to be the only known replication competent member both in vitro and in vivo (Laderoute et al., 2007; Laderoute et al., 2015). Accordingly, since phylogenetic evidence supports an association of HERV-K HML-2 activity with protection against integration of orthoretroviruses (i.e., HIV-1 acquisition), this may help substantiate the notion that HERV-K102 particles, along with expression of proteins from other HML-2 elements, might help prevent HIV-1 acquisition and provide sterilizing immunity (Image 1).
Image 1. Evidence that HERV-K102 Replicative Activity (increased genomic integration) Is Associated with Sterilizing Immunity Against HIV-1 Acquisition in an HIV-1 Exposed Seronegative (HESN) Cohort of Commercial Sex Workers (CSW). Extreme Protection Against Pandemic Viruses.
On the curious origins of HERV-K102 in humans
Somewhat ironically, humans apparently acquired the HERV-K102 defense mechanism from the same source of the modern HIV-1 pandemic strain; namely, chimpanzees, possibly between 500,000 and up to 2 mya (Romano et al., 2006; Subramanian et al., 2011). The Homo-Pan split has been estimated at 6.6 mya (Magiorkinis et al., 2015) or earlier at 7-8 mya (Langergraber et al., 2012).
As mentioned, the HERV-K HML-2 elements originated in primates about 10.3 mya and the CERV-K102 sequence (DQ112149), which is 97% identical to HERV-K102, was estimated to have integrated into chimpanzees at a non-orthologous position about 10 (+/- 3.3) Mya (Romano et al., 2006).
Lentiviruses like HIV-1 may have been active in primates since the divergence of chimpanzees and humans (Katzourakis et al., 2007; Sawyer et al., 2004). Moreover, it has been suggested the ancestor to HIV-1 may have arisen in chimpanzees about 4 mya (Gifford, 2012). Since it has been reported that subsets of chimpanzees with chronic HIV-1 infection showed progression analogous to humans, including greater expression of CD38 in CD8+ HLADR+ T cells (O’Neil et al., 2000), this raises the notion that an HERV-K102 ancestor, as a potential antidote for HIV-1 infection may have been selected through evolution in chimpanzees before it was acquired by humans. The genus Homo arose about 2 mya (Khan N et al., Genome Biol Evol, 2020). Accordingly, it is possible over about a 2 million-year window or longer, the HERV-K102 ancestor may have adapted to an HIV-1 like ancestor lentivirus(es) in chimpanzees prior to its acquisition by humans. Indeed, there is genetic evidence from a 5 amino acid deletion fixed in a human orthologue of tetherin, that humans may have been afflicted by a lentivirus presence long before HIV-1, perhaps about 800,000 years ago, which may have caused changes in innate immune genes in humans (Compton AA et al., Philos Trans R Soc Lond B Biol Sci. 2013). Thus, the phylogenetic evidence raises the notion that HERV-K102 as a replication competent HERV-K HML-2 retrovirus, may have evolved specific mechanisms to limit HIV/lentivirus replication and retrovirus genome invasion in chimpanzees before being acquired in humans.
Extinct hominins lost HERV-K102 at the orthologous position at 1q22
It is important to note that there is also additional phylogenetic evidence consistent with HERV-K102 protection generally against emerging RNA viruses, and not necessarily limited to orthoretroviruses.
Neanderthals and Denisovans, appear to have lost HERV-K102 at the orthologous chromosome position at 1q22 (Image 2.) and both went extinct. The timing of the last common ancestor was about 0.6 million years ago whereas the Homo species which includes Neanderthals and Denisovans, arose about 2 million years ago (Khan N et al., Genome Biol Evol, 2020). As deletions may be common at the time of insertions, this data raises the possibility that HERV-K102 may have been acquired from chimpanzees by the last common hominin ancestor about 0.6 mya. It should be noted that the earliest modern human remains in Europe so far discovered was about 45,000 years ago (Hajdinjak M et al., Nature, 2021). Thus, at the time of HERV-K102 acquisition the common ancestor would have resided in Africa and this makes it plausible that HERV-K102 may have been acquired from chimpanzees known to live in Africa but not Europe. The finding also by Compton et al. (2013) referenced above implicates an exposure to chimpanzee lentiviruses around the same time, consistent with co-mingling of humans and chimpanzees at that time that HERV-K102 crossed over and into humans.
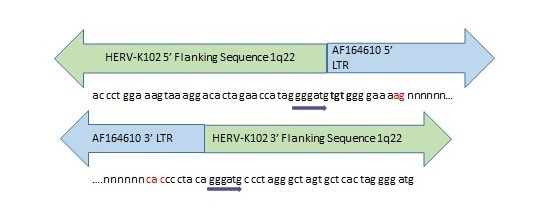
Image 2. Evidence for Past Integration and Excision of HERV-K102 at Orthologous Positions in the Neanderthal and Denisovan Genomes.
Direct repeats (gggatg) flank the orthologous HERV-K102 sequence in the human and extinct hominin genomes. The nucleotides marked in red were missing from the Denisovan orthologous position but present in the Neanderthal genome. Most of the HERV-K102 provirus was missing in both extinct hominins with only a few nucleotides corresponding to the ends of the LTRs remaining intact. “n” stands for missing nucleotides. Inquiry was made of the Altai Neanderthal or Denisovan genome at http://bioinf.eva.mpg.de/fetchseq/ on chromosome 1 strand at 155,596,423 to 155,605,644 by permission from Dr. Svante Paabo. AF164610 GenBank LTR flanking sequences used: 5’ LTR sequence = AF095801 and 3’ LTR sequence = AF095802.
RNA virus pandemics likely played some role in the demise of the extinct hominins as adaptively introgressed Neanderthal genes in humans frequently feature innate immunity genes against RNA viruses (Enard D & Petro DA, Cell, 2018) or immune pathways against pathogens (Gouy A & Excoffier L, Mol Biol Evol, 2020). As well, innate immune reactions with pathogens affected human evolution between 0.6 and 2 mya (Khan N et al., Genome Biol Evol, 2020).
Consistent with this notion of a superior innate defense system of humans versus other interacting hominins was the finding of successful admixture between humans and Neanderthals but only where the female was human. This was deduced by showing mitochondrial DNA carried from the egg (ie., female) in humans contained no introgressed Neanderthal genes (discussed in Bucking R et al., BMC Genomics, 2019). Accordingly this may further substantiate the extreme protection the HERV-K102 system offers against novel epidemics.
However, it should be appreciated that there may have been other reasons for this one-way admixture aside from innate immunity and/or that caused the extinction of the Neanderthals and Denisovans.
Nevertheless, it is tempting to speculate that at least in part, the HERV-K102 protection system may have contributed to the survival of Homo sapiens.
If so, one can immediately appreciate the significance of the HERV-K102 protector system against a deadly RNA pandemic virus like SARS-CoV-2, or sterilizing immunity against HIV-1.
REFERENCES
Bücking R, Cox MP, Hudjashov G, Saag L, Sudoyo H, Stoneking M. Archaic mitochondrial DNA inserts in modern day nuclear genomes. BMC Genomics. 2019 Dec 26;20(1):1017. doi: 10.1186/s12864-019-6392-8.
Chuong EB, Elde NC, Feschotte C. Regulatory evolution of innate immunity through co-option of endogenous retroviruses. Science. 2016 Mar 4;351(6277):1083-7. doi: 10.1126/science.aad5497.
Compton AA, Malik HS, Emerman M. Host gene evolution traces the evolutionary history of ancient primate lentiviruses. Philos Trans R Soc Lond B Biol Sci. 2013 Aug 12;368(1626):20120496. doi: 10.1098/rstb.2012.0496.
Enard D, Petrov DA. Evidence that RNA Viruses Drove Adaptive Introgression between Neanderthals and Modern Humans. Cell. 2018 Oct 4;175(2):360-371.e13. doi: 10.1016/j.cell.2018.08.034.
Gifford RJ. Viral evolution in deep time: lentiviruses and mammals. Trends Genet. 2012 Feb;28(2):89-100. doi: 10.1016/j.tig.2011.11.003.
Gouy A, Excoffier L. Polygenic Patterns of Adaptive Introgression in Modern Humans Are Mainly Shaped by Response to Pathogens. Mol Biol Evol. 2020 May 1;37(5):1420-1433. doi: 10.1093/molbev/msz306. Hajdinjak M 2021 Nature
Katzourakis A, Tristem M, Pybus OG, Gifford RJ. Discovery and analysis of the first endogenous lentivirus. Proc Natl Acad Sci U S A. 2007 Apr 10;104(15):6261-5. doi: 10.1073/pnas.0700471104.
Khan N, de Manuel M, Peyregne S, Do R, Prufer K, Marques-Bonet T, Varki N, Gagneux P, Varki A. Multiple Genomic Events Altering Hominin SIGLEC Biology and Innate Immunity Predated the Common Ancestor of Humans and Archaic Hominins. Genome Biol Evol. 2020 Jul 1;12(7):1040-1050. doi: 10.1093/gbe/evaa125.
Kim YJ, Han K. Endogenous retrovirus-mediated genomic variations in chimpanzees. Mob Genet Elements. 2015 Feb 3;4(6):1-4. doi: 10.4161/2159256X.2014.990792.
Laderoute MP, Giulivi A, Larocque L, Bellfoy D, Hou Y, Wu HX, Fowke K, Wu J, Diaz-Mitoma F. The replicative activity of human endogenous retrovirus K102 (HERV-K102) with HIV viremia. AIDS. 2007 Nov 30;21(18):2417-24. doi: 10.1097/QAD.0b013e3282f14d64.
Laderoute MP, Larocque LJ, Giulivi A, Diaz-Mitoma F. Further Evidence that Human Endogenous Retrovirus K102 is a Replication Competent Foamy Virus that may Antagonize HIV-1 Replication. Open AIDS J. 2015 Dec 7;9:112-22. doi: 10.2174/1874613601509010112.
Langergraber KE, Prüfer K, Rowney C, Boesch C, Crockford C, Fawcett K, Inoue E, Inoue-Muruyama M, Mitani JC, Muller MN, Robbins MM, Schubert G, Stoinski TS, Viola B, Watts D, Wittig RM, Wrangham RW, Zuberbühler K, Pääbo S, Vigilant L. Generation times in wild chimpanzees and gorillas suggest earlier divergence times in great ape and human evolution. Proc Natl Acad Sci U S A. 2012 Sep 25;109(39):15716-21. doi: 10.1073/pnas.1211740109.
Magiorkinis G, Blanco-Melo D, Belshaw R. The decline of human endogenous retroviruses: extinction and survival. Retrovirology. 2015 Feb 2;12:8. doi: 10.1186/s12977-015-0136-x.O’Neill 2000
Romano CM, Ramalho RF, Zanotto PM. Tempo and mode of ERV-K evolution in human and chimpanzee genomes. Arch Virol. 2006 Nov;151(11):2215-28. doi: 10.1007/s00705-006-0792-1.
Sawyer SL, Emerman M, Malik HS. Ancient adaptive evolution of the primate antiviral DNA-editing enzyme APOBEC3G. PLoS Biol. 2004 Sep;2(9):E275. doi: 10.1371/journal.pbio.0020275.
Subramanian RP, Wildschutte JH, Russo C, Coffin JM. Identification, characterization, and comparative genomic distribution of the HERV-K (HML-2) group of human endogenous retroviruses. Retrovirology. 2011 Nov 8;8:90. doi: 10.1186/1742-4690-8-90.






FASCINATING!!!
Thank you.